Home | Research Accomplishments | Services & Requests | Equipment
To request services please visit the MSR iLab page.
Metabolomics is quantitative characterization of metabolites. The metabolome represents the collection of all metabolites in a biological system, such as cell, tissue, organ, or organism. Metabolites are not only substrates and products of biochemical reactions, many of them also serve as regulators of enzymatic activities. Metabolite levels reflect the balance between production and consumption. Stable-isotope tracers allow more direct measurement of metabolite production rates, i.e., pathway activities (“fluxes”). The combination of concentration and flux measurement provides a more complete view of metabolism and its role in cancer.

To help our investigators benefit from these new methods and technologies, we provide the following services:
Water-Soluble Metabolite Quantitation
Our most popular service. Covers classical intermediary metabolites including glycolytic intermediates, TCA compounds, amino acids, nucleotides, and derivatives. Specificity is achieved through the combination of chromatographic separation and high-resolution MS, with MS/MS used in selected cases to differentiate isomers. Current methods cover approximately 300 water-soluble metabolites, each identified by retention time match to synthetic standards. For researchers primarily interested in central carbon metabolism, it is common to analyze only anionic metabolites (carboxylic acids and phosphorylated compounds), which can be achieved with a single LC-MS run in negative ion mode. To also cover cationic metabolites like acyl-carnitines, nucleosides, and polyamines, users may elect to also run in positive ion mode. Stable isotope labeling patterns can also be routinely measured (13C, 15N, 2H, etc.).
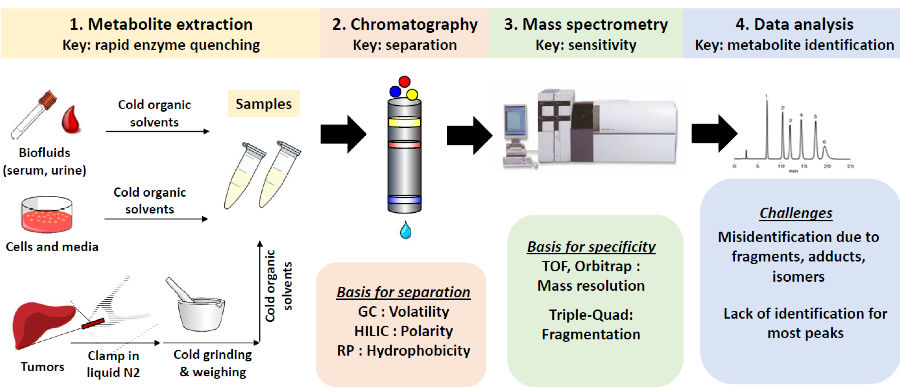
Figure. Metabolomics workflow
Lipid Quantitation
Identification and quantitation of non-polar metabolites by LC-MS, with the coverage of approximately 500 lipids spanning all major classes. For isotope tracer studies, due to the challenge in extracting labeling data across such a complex mixture of lipids, users typically focus on saponified fatty acids. The labeling pattern of fatty acids provides high-value information regarding the metabolic sources of their building blocks: acetyl-CoA and NADPH.
Spatial Metabolomics
We are establishing a spatial metabolomics service based on high-resolution MALDI-imaging mass spectrometry using a Bruker SOLARIX instrument, with 30 μm spatial resolution, capable of distinguishing tumor regions (e.g., hypoxic core versus proliferative perimeter) but not single cells. To support the development of single-cell resolution using imaging mass spectrometry, we tested a Thermo SMALDI-Orbitrap system and are installing and testing a Bruker timsTOF fleX MALDI-2 with similar capabilities.
Novel or Unexpected Metabolite Discovery
The analytical methods used for metabolite quantitation are untargeted: They measure all ions present in the sample. From these measurements, we routinely quantitate only known analytes, because most users are focused on the abundances and labeling of these classical species (like glutamine, ATP, fructose bisphosphate). While there are a very large number of other peaks, most reflect analytical artifacts, not metabolites. Nevertheless, the discovery of new cancer-associated metabolites is a great scientific opportunity, born out by the clinical importance of 2-hydroxyglutarate. Online tools (XCMS, MzMine2) and commercial software (CompoundDiscoverer) enable users to find unexpected peaks that go up or down in a particular set of samples. The shared resource then works with users to identify the underlying metabolites. To this end, the Metabolomics shared resource is developing novel strategies for metabolite identification based on combining isotope labeling and MS/MS.
Expert Consultation and Training
Investigators planning to conduct research using the shared resource are encouraged to submit an experimental plan describing the biological question and protocol design. Users are guided to detailed SOPs for critical steps like sample harvesting and metabolite extraction. Given the unique requirements of many projects, Metabolomics personnel are available to meet with investigators at the inception of the project. Consultation regarding experimental design is also routinely provided prior to grant submission. After sample analysis, experimental data is provided to the investigator along with suggestions for interpretation. Additional experiments may be planned with the investigator to optimize the results. In addition, the Metabolomics shared resource offers seminars and presentations by internal and external experts. It trains users in the Metabolomics Analysis and Visualization Engine (MAVEN) software package for data processing and visualization. To facilitate the training of graduate students and postdocs, professional staff manually review the LC-MS raw data of less experienced users, to ensure accurate processing and interpretation. Facility staff members also routinely manually check all data associated with key biological discoveries.
Last updated 01/23/2023

