Home | Research Accomplishments | Services & Requests | Equipment | BISR Utilization Voucher Application Form
Biomedical Imaging
TMA-Analysis Workstation
High-res digital camera with RGB liquid crystal filter (Olympus DP70 and DP71) to digitize specimens
Computer-assisted microscopy (Olympus AX70 and BX40 with six-way robotic stage/motorized turret) to support telemicroscopy and telepathology applications
Informatica-based extraction transformation and load interface (ETL) to automatically populate a secure, IRB-approved, Oracle-based Clinical and Research Data Warehouse with data elements originating from the multi-modal data sources. During empirical benchmark experiments this interface was able to reliably process and load all of the associated data for 5,000 breast cancer cases into the Warehouse in ~12 minutes
Nuance multispectral imaging system for accurate spectral classification and unmixing. It can be configured for either bright-field or fluorescence microscopy in the visible (VIS) wavelength range of 420-720nm (up to 950nm for fluorescence-only systems) to allow systematic investigation regarding the use of metamerism, manifold shortcuts and digital staining applications

Multi-spectral Imaging Workstation
Nikon Eclipse 90i fully automated bright-field/fluorescent microscopic imaging station with Photometric Coolsnap ES2 camera to support quick, reliable subsampling of specimens for presentations and publications
Olympus VS120 bright-field/fluorescent whole slide scanner with 100 slide capacity and standalone processing workstation which generates high-resolution digital images from microscopy samples within minutes while optimizing scan speed, Z-positioning and efficiency
Image server and graphical interface within Rutgers firewall to allow secure review and annotation by consulting pathologists and collaborators or to allow quantitative analysis using a wide range of computational imaging tools that are made available through the shared resource
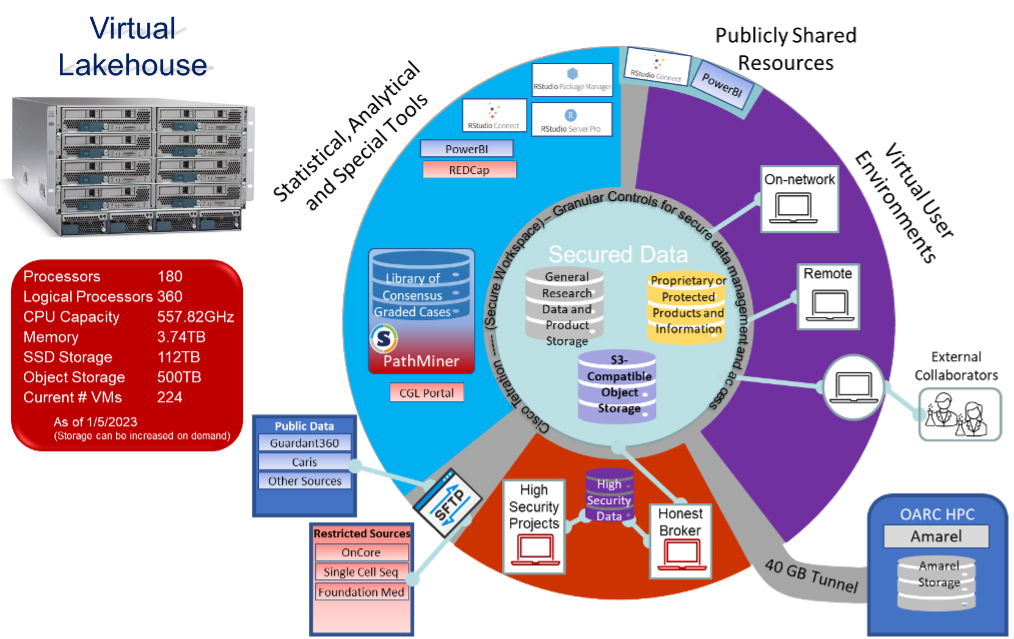
Lakehouse
In response to the COVID-19 pandemic, BISR worked with Rutgers Cancer Institute IT to respond to an increased demand for remote access and analysis of large data sets while simultaneously maintaining onsite capabilities. In this environment, BISR applications provide Windows and Linux based virtual machines to give researchers more effective tools and ability to access and process large data sets. This environment addresses security and computing needs in a flexible manner, providing centralized access to a robust selection of tools for statistical analysis, storage, file transfers, and direct access to our high-performance compute clusters. A sample of the tools that will be available in this environment are: Posit (formerly RStudio) Workspace and Posit Connect, sFTP, PathMiner, Amarel Cluster (HPC), Sandbox Testing Environment, REDCap, Microsoft Power BI Report Server, Flexible End User Virtual Desktops. The BISR works closely with Rutgers Cancer Institute IT to expand on a growing number of tools available to support our research and clinical applications.
Amarel
The Amarel cluster provides a shared platform that optimizes resources for the benefit of all users. Named in honor of Dr. Saul Amarel, one of the founders of the Rutgers Computer Science Department and contributor to advanced computing and artificial intelligence research and methodologies, Amarel is designed to suit many different research applications.
- Traditional high-performance computing
- High Throughput Computing (HTC)
- Data Analytics
- Large memory systems
- Virtualization
- Accelerators (GPGPUs, XEON PHIs, FPGAs)
- The ability to couple with edge devices
Using a condo model, Amarel provides 700 compute nodes with 20,980 Intel Xeon cores, 190 GPUs, Open OnDemand servers and InfiniBand FDR and EDR fabric. Within this condo model, Rutgers Cancer Institute owns 11 nodes that can be dedicated to our specific needs and we have access to the remaining nodes on an as available basis.
Updated 02/01/2023

