Home | Research Accomplishments | Services & Requests | Equipment
To request services, please contact the CGSR Director, Patricia Soteropoulos. Services will be transfer to iLab in early 2023. Also starting in 2023, a new price subsidy of 15% will be implemented for Rutgers Cancer Institute members assigned full membership status.
The CGSR works closely with basic and clinical researchers to provide full support for experimental design through data analysis. The array of services includes making recommendations on technology platforms and workflows, performing sample processing, and providing support for data analysis, data interpretation, grant writing, and manuscript preparation. Rutgers Cancer Institute member projects are directed to the Bioinformatics SR for bioinformatics analysis.
The CLIA-certified, CAP-accredited clinical genomics laboratory offers cytogenetic tests, including fluorescent in situ hybridization (FISH) on peripheral blood, bone marrow, lymph node, solid tumors, and paraffin-embedded tissue and genetic testing for chromosomal abnormalities associated with leukemias, lymphomas, and various solid tumors including head and neck and breast cancers.
High throughput sequencing and microarray platforms include Illumina MiniSeq, MiSeq, NextSeq, and NovaSeq sequencers, an Oxford Nanopore GridION system, Affymetrix GeneChips, and Agilent Microarrays. In addition, the Center works with other regional core facilities to provide access to PacBio sequencers.
All associated equipment for sample processing and quality control, including an Agilent BioAnalyzer, Agilent TapeStation, NanoDrop, and Qubit, are also available. Sanger sequencing and fragment analysis are performed on the Applied Biosystems SeqStudio Genetic Analyzer.
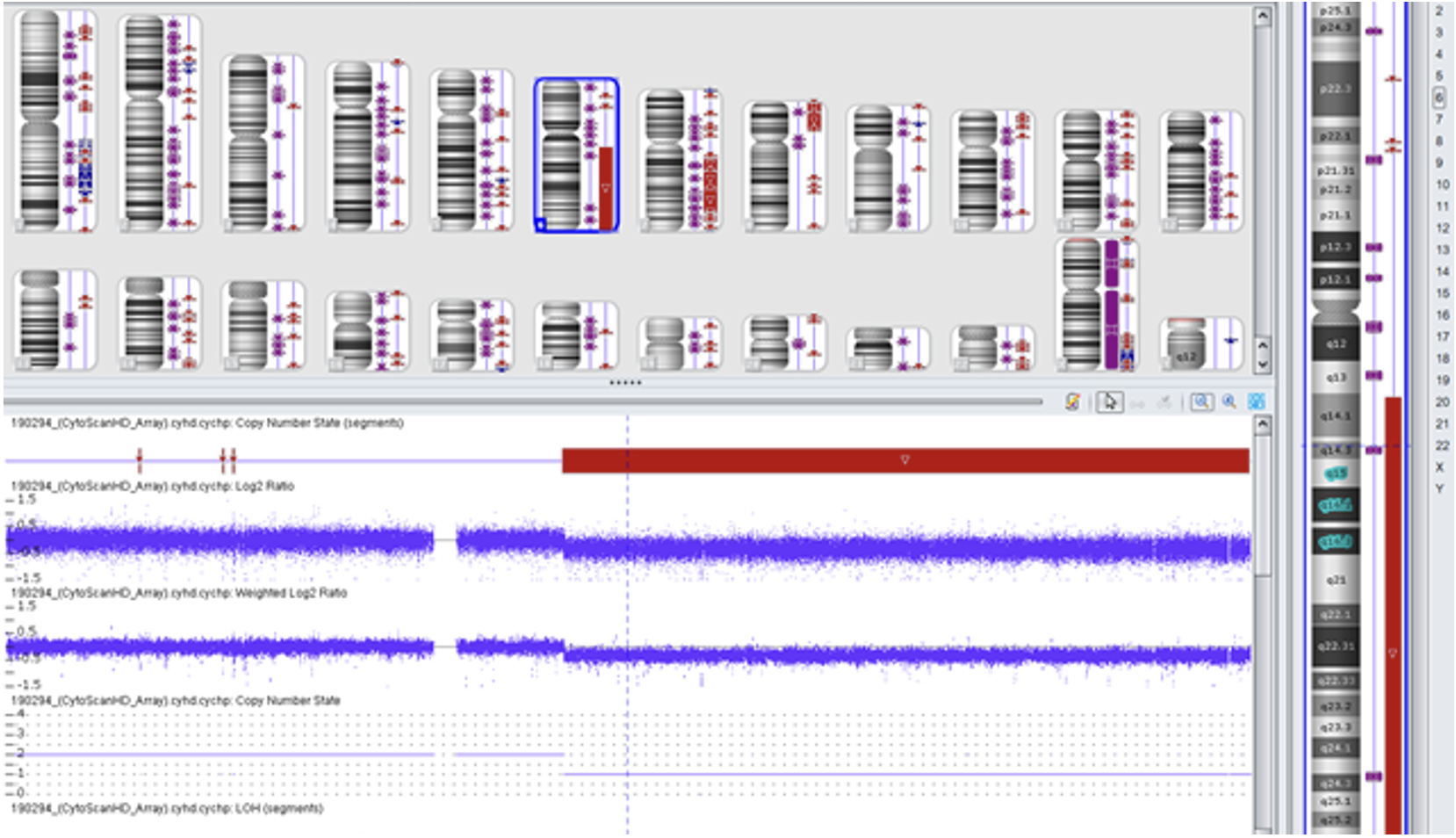
Example of cytogenetics services offered:
Last updated 01/20/2023

